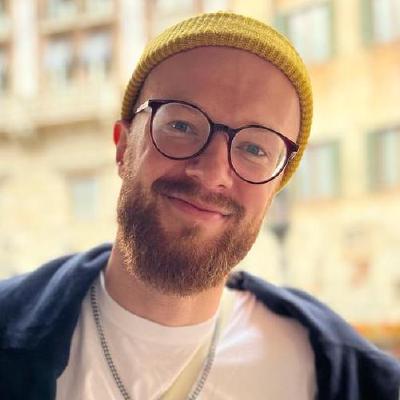
Introducing mle-monitor: A Lightweight Experiment & Resource Monitoring Tool 📺
Published:
“Did I already run this experiment before? How many resources are currently available on my cluster?” If these are common questions you encounter during your daily life as a researcher, then mle-monitor is made for you. It provides a lightweight API for tracking your experiments using a pickle protocol database (e.g. for hyperparameter searches and/or multi-configuration/multi-seed runs). Furthermore, it comes with built-in resource monitoring on Slurm/Grid Engine clusters and local machines/servers. Finally, it leverages rich in order to provide a terminal dashboard that is updated online with new protocolled experiments and the current state of resource utilization. Here is an example of a dashboard running on a Grid Engine cluster:

%load_ext autoreload
%autoreload 2
%config InlineBackend.figure_format = 'retina'
try:
import mle_monitor
except:
!pip install -q mle-monitor
import mle_monitor
mle-monitor comes with three core functionalities:
MLEProtocol: A composable protocol database API for ML experiments.MLEResource: A tool for obtaining server/cluster usage statistics.MLEDashboard: A dashboard visualizing resource usage & experiment protocol.
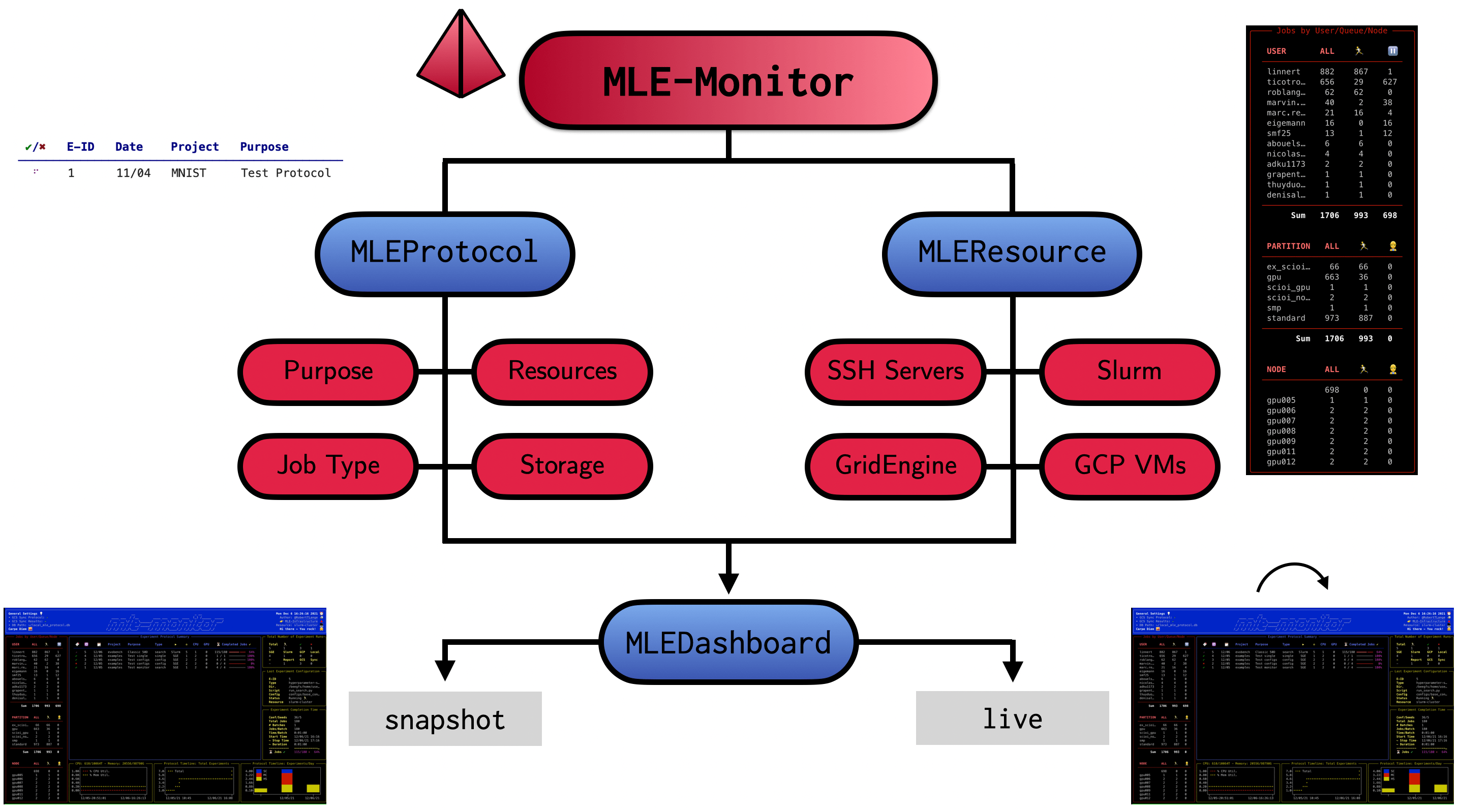
Finally, mle-monitor is part of the mle-infrastructure and comes with a set of handy built-in synergies. We will wrap-up by outlining a full workflow of using the protocol together with a random search experiment using mle-hyperopt, mle-scheduler and mle-logging.
# Check if code is run in Colab: If so -- download configs from repo
try:
import google.colab
IN_COLAB = True
!wget -q https://raw.githubusercontent.com/mle-infrastructure/mle-monitor/main/examples/train.py
!wget -q https://raw.githubusercontent.com/mle-infrastructure/mle-monitor/main/examples/base_config.json
except:
IN_COLAB = False
Experiment Management with MLEProtocol 📝
from mle_monitor import MLEProtocol
# Load the protocol from a local file (create new if it doesn't exist yet)
protocol = MLEProtocol(protocol_fname="mle_protocol.db", verbose=True)
In order to add a new experiment to the protocol database you have to provide a dictionary containing the experiment meta data:
| Search Type | Description | Default |
|---|---|---|
purpose |
Purpose of experiment | 'None provided' |
project_name |
Project name of experiment | 'default' |
exec_resource |
Resource jobs are run on | 'local' |
experiment_dir |
Experiment log storage directory | 'experiments' |
experiment_type |
Type of experiment to run | 'single' |
base_fname |
Main code script to execute | 'main.py' |
config_fname |
Config file path of experiment | 'base_config.yaml' |
num_seeds |
Number of evaluations seeds | 1 |
num_total_jobs |
Number of total jobs to run | 1 |
num_job_batches |
Number of jobs in single batch | 1 |
num_jobs_per_batch |
Number of sequential job batches | 1 |
time_per_job |
Expected duration: days-hours-minutes | '00:01:00' |
num_cpus |
Number of CPUs used in job | 1 |
num_gpus |
Number of GPUs used in job | 0 |
meta_data = {
"purpose": "Test Protocol", # Purpose of experiment
"project_name": "MNIST", # Project name of experiment
"exec_resource": "local", # Resource jobs are run on
"experiment_dir": "log_dir", # Experiment log storage directory
"experiment_type": "hyperparameter-search", # Type of experiment to run
"base_fname": "train.py", # Main code script to execute
"config_fname": "base_config.json", # Config file path of experiment
"num_seeds": 5, # Number of evaluations seeds
"num_total_jobs": 10, # Number of total jobs to run
"num_jobs_per_batch": 5, # Number of jobs in single batch
"num_job_batches": 2, # Number of sequential job batches
"time_per_job": "00:05:00", # Expected duration: days-hours-minutes
"num_cpus": 2, # Number of CPUs used in job
"num_gpus": 1, # Number of GPUs used in job
}
e_id = protocol.add(meta_data, save=False)
[15:37:51] INFO Added experiment 1 to protocol. mle_protocol.py:162
Adding the experiment will load in the configuration file (either .json or .yaml) and set the experiment status to “running”. You can then always retrieve the provided information using protocol.get(e_id):
protocol.get(e_id)
{'purpose': 'Test Protocol',
'project_name': 'MNIST',
'exec_resource': 'local',
'experiment_dir': 'log_dir',
'experiment_type': 'hyperparameter-search',
'base_fname': 'train.py',
'config_fname': 'base_config.json',
'num_seeds': 5,
'num_total_jobs': 10,
'num_jobs_per_batch': 5,
'num_job_batches': 2,
'time_per_job': '00:05:00',
'num_cpus': 2,
'num_gpus': 1,
'git_hash': '60cb3e3883da3888865b47abf0d5b6257e6d91e5',
'loaded_config': [{'train_config': {'lrate': 0.1},
'model_config': {'num_layers': 5},
'log_config': {'time_to_track': ['step_counter'],
'what_to_track': ['loss'],
'time_to_print': ['step_counter'],
'what_to_print': ['loss'],
'print_every_k_updates': 10,
'overwrite_experiment_dir': 1}}],
'e-hash': '897b974332747d81e84b3ed688f7862d',
'retrieved_results': False,
'stored_in_cloud': False,
'report_generated': False,
'job_status': 'running',
'completed_jobs': 0,
'start_time': '12/09/21 15:37',
'duration': '0:10:00',
'stop_time': '12/10/21 01:37'}
You can also always print a summary snapshot of the last experiments using protocol.summary(). By providing the boolean option full, you also print the resources used in an experiment:
# Print a summary of the last experiments
sub_df = protocol.summary()
# ... and a more detailed version
sub_df = protocol.summary(full=True)
🔖 🆔 🗓 Project Purpose Type ▶ ♻ CPU GPU ──────────────────────────────────────────────────────────────────────────── ⠸ 1 12/09 MNIST Test Protocol search Local 5 2 1
🔖 🆔 🗓 Project Purpose Type ▶ ♻ CPU GPU ⏳ Completed Jobs ✔ ─────────────────────────────────────────────────────────────────────────────────────────── ⠼ 1 12/09 MNIST Test Protocol search Local 5 2 1 0 /10 0%
If you want to ad-hoc change any of the stored attributes of an experiment, you can do so using the update method. Furthermore, you can change the experiment status using abort or complete:
# Update some element in the database
protocol.update(e_id, "exec_resource", "slurm-cluster", save=False)
# Abort the experiment - changes status
protocol.abort(e_id, save=False)
sub_df = protocol.summary()
# Get the status of the experiment
protocol.status(e_id)
🔖 🆔 🗓 Project Purpose Type ▶ ♻ CPU GPU ──────────────────────────────────────────────────────────────────────────── ✖ 1 12/09 MNIST Test Protocol search Slurm 5 2 1
'aborted'
If you would like to get a summary of all reported experiments, the last experiment and its resource requirements, protocol.monitor() does so:
# Get the monitoring data - used later in dashboard
protocol_data = protocol.monitor()
protocol_data["total_data"]
{'total': '1',
'run': '0',
'done': '0',
'aborted': '1',
'sge': '0',
'slurm': '1',
'gcp': '0',
'local': '0',
'report_gen': '0',
'gcs_stored': '0',
'retrieved': '0'}
Finally, you can also store other data specific to an experiment using an additional dictionary of data as follows:
extra_data = {"extra_config": {"lrate": 3e-04}}
e_id = protocol.add(meta_data, extra_data, save=False)
protocol.get(e_id)["extra_config"]
[15:37:58] INFO Added experiment 2 to protocol. mle_protocol.py:162
{'lrate': 0.0003}
Syncing your Protocol DB with a GCS Bucket
If you would like to keep a remote copy of your protocol, you can also automatically sync your protocol database with a Google Cloud Storage (GCS) bucket. This is especially useful when running experiments on multiple resource and will require you to have created a GCP project and a GCS bucket. Furthermore you will have to provide you .json authentication key path. If you don’t have one yet, have a look here. Alternatively, just make sure that the environment variable GOOGLE_APPLICATION_CREDENTIALS is set to the right path.
# Sync your protocol with a GCS bucket
cloud_settings = {
"project_name": "mle-toolbox", # Name of your GCP project
"bucket_name": "mle-protocol", # Name of your GCS bucket
"protocol_fname": "mle_protocol.db", # Name of DB file in GCS bucket
"use_protocol_sync": True, # Whether to sync the protocol
"use_results_storage": False # Whether to upload zipped dir at completion
}
protocol = MLEProtocol(protocol_fname="mle_protocol.db",
cloud_settings=cloud_settings,
verbose=True)
[15:38:01] INFO No DB found in GCloud Storage - mle_protocol.db gcs_sync.py:39
INFO New DB will be created - mle-toolbox/mle-protocol gcs_sync.py:40
INFO Pulled protocol from GCS bucket: mle-protocol. mle_protocol.py:379
e_id = protocol.add(meta_data)
[15:38:06] INFO Added experiment 1 to protocol. mle_protocol.py:162
INFO Locally stored protocol: mle_protocol.db mle_protocol.py:91
[15:38:07] INFO Send to GCloud Storage - mle_protocol.db gcs_sync.py:70
INFO Send protocol to GCS bucket: mle-protocol. mle_protocol.py:364
INFO GCS synced protocol: mle_protocol.db mle_protocol.py:97
Finally, you can also choose to store the results of an experiment in the GCS bucket. In this case the protocol will upload a zipped version of your created experiment_dir to the bucket whenever you call protocol.complete().
Resource Monitoring with MLEResource 📉
You can monitor your local machine, server or clusters using the MLEResource. If you are running this on a Google Colab, make sure to add a GPU accelerator!
from mle_monitor import MLEResource
resource = MLEResource(resource_name="local")
resource_data = resource.monitor()
resource_data["user_data"].keys()
dict_keys(['pid', 'p_name', 'mem_util', 'cpu_util', 'cmdline', 'total_cpu_util', 'total_mem_util'])
You can also monitor slurm or grid engine clusters by providing the queues/partitions to monitor in monitor_config:
resource = MLEResource(
resource_name="slurm-cluster",
monitor_config={"partitions": ["<partition-1>", "<partition-2>"]},
)
resource = MLEResource(
resource_name="sge-cluster",
monitor_config={"queues": ["<queue-1>", "<queue-2>"]}
)
Dashboard Visualization with MLEDashboard 🎞️
from mle_monitor import MLEDashboard
dashboard = MLEDashboard(protocol, resource)
# Get a static snapshot of the protocol & resource utilisation
# Note: This will look a lot nicer in your terminal!
dashboard.snapshot()
# Run monitoring in while loop - dashboard
dashboard.live()
- Add widget animation!/screenshot
Integration with the MLE-Infrastructure Ecosystem 🔺
Running a Hyperparameter Search for Multiple Random Seeds
try:
from mle_hyperopt import RandomSearch
from mle_scheduler import MLEQueue
from mle_logging import load_meta_log
except:
!pip install -q mle-hyperopt mle-scheduler mle-logging
!pip install --upgrade rich
from mle_hyperopt import RandomSearch
from mle_scheduler import MLEQueue
from mle_logging import load_meta_log
We again start by adding an experiment to the protocol at launch time.
# Load (existing) protocol database and add experiment data
protocol_db = MLEProtocol("mle_protocol.db", verbose=True)
meta_data = {
"purpose": "random search", # Purpose of experiment
"project_name": "surrogate", # Project name of experiment
"exec_resource": "local", # Resource jobs are run on
"experiment_dir": "logs_search", # Experiment log storage directory
"experiment_type": "hyperparameter-search", # Type of experiment to run
"base_fname": "train.py", # Main code script to execute
"config_fname": "base_config.json", # Config file path of experiment
"num_seeds": 2, # Number of evaluations seeds
"num_total_jobs": 4, # Number of total jobs to run
"num_jobs_per_batch": 4, # Number of jobs in single batch
"num_job_batches": 1, # Number of sequential job batches
"time_per_job": "00:00:02", # Expected duration: days-hours-minutes
}
new_experiment_id = protocol_db.add(meta_data)
[15:38:26] INFO Added experiment 2 to protocol. mle_protocol.py:162
INFO Locally stored protocol: mle_protocol.db mle_protocol.py:91
Afterwards, we leverage mle-hyperopt to instantiate a random search strategy with its parameter space. We then ask for two configurations and store them as .yaml files in our working directory:
# Instantiate random search class
strategy = RandomSearch(
real={"lrate": {"begin": 0.1, "end": 0.5, "prior": "log-uniform"}},
integer={"batch_size": {"begin": 1, "end": 5, "prior": "uniform"}},
categorical={"arch": ["mlp", "cnn"]},
verbose=True,
)
# Ask for configurations to evaluate & run parallel eval of seeds * configs
configs, config_fnames = strategy.ask(2, store=True)
configs
MLE-Hyperopt Random Search Hyperspace 🚀 🌻 Variable Type Search Range ↔ ────────────────────────────────────────────────────────────────────── arch categorical ['mlp', 'cnn'] lrate real Begin: 0.1, End: 0.5, Prior: log-uniform batch_size integer Begin: 1, End: 5, Prior: uniform
[{'arch': 'mlp', 'lrate': 0.360379148648584, 'batch_size': 3},
{'arch': 'cnn', 'lrate': 0.26208630215377515, 'batch_size': 2}]
Next, we can use a MLEQueue from mle-scheduler to run our training script train.py for our two configurations and two different random seeds. train.py implements a simple surrogate training loop, which logs some statistics with the help of mle-logging. Afterwards, we merge the resulting logs into a single meta_log.hdf5 and retrieve the mean (over seeds) test loss score for both configurations.
queue = MLEQueue(
resource_to_run="local",
job_filename="train.py",
config_filenames=config_fnames,
random_seeds=[1, 2],
experiment_dir="logs_search",
protocol_db=protocol_db,
)
queue.run()
# Merge logs of random seeds & configs -> load & get final scores
queue.merge_configs(merge_seeds=True)
meta_log = load_meta_log("logs_search/meta_log.hdf5")
test_scores = [meta_log[r].stats.test_loss.mean[-1] for r in queue.mle_run_ids]
Output()
[15:38:35] INFO Locally stored protocol: mle_protocol.db mle_protocol.py:91
INFO Locally stored protocol: mle_protocol.db mle_protocol.py:91
INFO Locally stored protocol: mle_protocol.db mle_protocol.py:91
[15:38:36] INFO Locally stored protocol: mle_protocol.db mle_protocol.py:91
MLEQueue - local • 4/4 Jobs ━━━━━━━━━━━━━━━━━━━━━━━━━━━ 100% • 0:00:01 ⌛
Finally, we update the random search strategy and tell the protocol that the experiment has been completed:
# Update the hyperparameter search strategy
strategy.tell(configs, test_scores)
# Wrap up experiment (store completion time, etc.)
protocol_db.complete(new_experiment_id)
┏━━━━━━━━━━━━━━━┳━━━━┳━━━━━━━━━┳━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━┓ ┃ 📥 Total: 2 ┃ ID ┃ Obj. 📉 ┃ Configuration 🔖 - 12/09/2021 15:38:43 ┃ ┡━━━━━━━━━━━━━━━╇━━━━╇━━━━━━━━━╇━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━━┩ │ Best Overall │ 0 │ 1.193 │ 'arch': 'mlp', 'lrate': 0.360379148648584, │ │ │ │ │ 'batch_size': 3 │ │ Best in Batch │ 0 │ 1.193 │ 'arch': 'mlp', 'lrate': 0.360379148648584, │ │ │ │ │ 'batch_size': 3 │ └───────────────┴────┴─────────┴───────────────────────────────────────────────┘
[15:38:43] INFO Locally stored protocol: mle_protocol.db mle_protocol.py:91
INFO Updated protocol - COMPLETED: 2 mle_protocol.py:253
Give it a try and let me know what you think! If you find a bug or are missing your favourite feature, feel free to contact me @RobertTLange or create an issue!

